Case Study in R: Recruitment agency
Last updated on January 19, 2026
1 About trees
1.1 Introduction
Applications
- Predict the land use of a given area (agriculture, forest, etc.) given satellite information, meteorological data, socio-economic information, prices information, etc.
- Determine how computer performance is related to a number of variables which describe the features of a PC (the size of the cache, the cycle time of the computer, the memory size and the number of channels. Both the last two were not measured but minimum and maximum values obtained).
Risk of heart attack
Predict high risk for heart attack:
- University of California: a study into patients after admission for a heart attack.
- 19 variables collected during the first 24 hours for 215 patients (for those who survived the 24 hours)
- Question: Can the high risk (will not survive 30 days) patients be identified?
A recruiting agency
Code
| Id | Dip | Test | Exp | Res |
|---|---|---|---|---|
| A | 1 | 5 | 4 | 0 |
| B | 2 | 3 | 3 | 0 |
| C | 1 | 4 | 5 | 1 |
| D | 2 | 3 | 4 | 0 |
| E | 1 | 4 | 4 | 0 |
| F | 4 | 3 | 4 | 1 |
| G | 3 | 4 | 4 | 1 |
| H | 1 | 1 | 5 | 0 |
| I | 3 | 2 | 5 | 1 |
| J | 5 | 4 | 4 | 1 |
Code
1.2 Full binary trees
1.3 When to use decision trees?
2 Procedure
2.1 Overview
A synthetic 2D example
Code
2.2 Recursive partitioning
Depth 0
Code
Depth 1
Code
Depth 2
Code
Depth 3
Code
2.3 Growing the tree
The need of an impurity criterion
Chosing an impurity criterion
We consider the split: \(X_1 <\) 1.64.
| X_1 | X_2 | Y | split |
|---|---|---|---|
| -1.5017872 | 1.6803619 | A | Left |
| -0.9233743 | -0.8534551 | A | Left |
| 1.5462761 | 3.3774228 | B | Left |
| -0.8320433 | 1.1174336 | A | Left |
| -1.0854224 | 2.2061932 | A | Left |
| 0.9922884 | 0.0171323 | A | Left |
| 2.6586580 | 0.3255937 | A | Right |
| -0.1726686 | -0.2422814 | A | Left |
| -0.8874201 | -0.9109120 | A | Left |
| -1.1729836 | -1.6521118 | A | Left |
| A | B | Sum | |
|---|---|---|---|
| Left | 766 | 63 | 829 |
| Right | 34 | 137 | 171 |
| Sum | 800 | 200 | 1000 |
| A | B | Sum | |
|---|---|---|---|
| Left | 0.9240048 | 0.0759952 | 1 |
| Right | 0.1988304 | 0.8011696 | 1 |
| Sum | 0.8000000 | 0.2000000 | 1 |
| A | B | Sum | Freq | Gini | |
|---|---|---|---|---|---|
| Left | 0.9240048 | 0.0759952 | 1 | 0.829 | 0.1404398 |
| Right | 0.1988304 | 0.8011696 | 1 | 0.171 | 0.3185938 |
| Sum | 0.8000000 | 0.2000000 | 1 | 1.000 | 0.3200000 |
The total difference in Gini index of this split is:
2.4 Stopping the tree
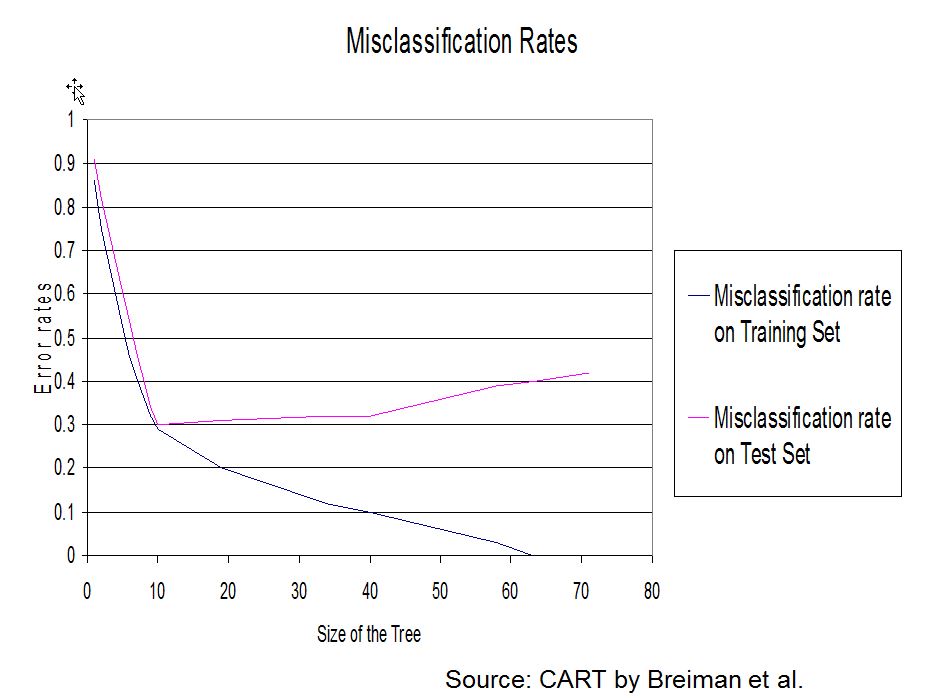
2.5 Pruning the tree
Why and how pruning?
A penalized criterion
Regression tree:
rpart(formula = Price ~ ., data = cu.summary, minbucket = 1,
cp = 0)
Variables actually used in tree construction:
[1] Country Mileage Reliability Type
Root node error: 7407472615/117 = 63311732
n= 117
CP nsplit rel error xerror xstd
1 2.5052e-01 0 1.00000 1.01330 0.15932
2 1.4836e-01 1 0.74948 0.86105 0.16322
3 8.7654e-02 2 0.60112 0.70608 0.14694
4 6.2818e-02 3 0.51347 0.58381 0.10526
5 3.4875e-02 4 0.45065 0.61862 0.12371
6 2.4396e-02 5 0.41577 0.61587 0.12360
7 1.1966e-02 8 0.34259 0.63034 0.12155
8 1.0640e-02 14 0.27079 0.68456 0.14321
9 9.9092e-03 15 0.26015 0.71913 0.14533
10 8.8587e-03 16 0.25024 0.72795 0.14614
11 7.3572e-03 20 0.21480 0.74085 0.15294
12 7.2574e-03 22 0.20009 0.73494 0.15289
13 3.8972e-03 28 0.15655 0.77054 0.15609
14 1.9968e-03 31 0.14334 0.78644 0.15729
15 1.9131e-03 33 0.13935 0.78699 0.15818
16 1.6070e-03 34 0.13744 0.78804 0.15816
17 1.1151e-03 35 0.13583 0.78647 0.15815
18 9.0617e-04 36 0.13471 0.79763 0.15711
19 4.7736e-04 42 0.12928 0.79600 0.15718
20 1.4084e-04 43 0.12880 0.79752 0.15718
21 1.0325e-04 45 0.12852 0.80091 0.15732
22 1.0187e-04 46 0.12841 0.80038 0.15734
23 8.0922e-05 47 0.12831 0.80051 0.15734
24 6.9751e-05 48 0.12823 0.80065 0.15733
25 6.0368e-05 49 0.12816 0.80068 0.15733
26 5.2584e-05 50 0.12810 0.80058 0.15733
27 5.1761e-05 52 0.12800 0.80058 0.15733
28 2.5252e-05 53 0.12794 0.79930 0.15735
29 1.2155e-05 54 0.12792 0.79968 0.15735
30 7.9655e-06 55 0.12791 0.79972 0.15734
31 1.5920e-06 56 0.12790 0.79958 0.15735
32 4.1013e-07 57 0.12790 0.79956 0.15735
33 0.0000e+00 58 0.12790 0.79958 0.15735Iterative pruning
Choosing the optimal subtree
3 Appendix
Call:
rpart(formula = Res ~ Dip + Test + Exp, data = candidates, method = "class",
parms = list(split = "gini"), control = rpart.control(minsplit = 2))
n= 10
CP nsplit rel error xerror xstd
1 0.80 0 1.0 2.0 0.0000000
2 0.10 1 0.2 0.2 0.1897367
3 0.01 3 0.0 0.6 0.2898275
Variable importance
Dip Test Exp
67 20 13
Node number 1: 10 observations, complexity param=0.8
predicted class=0 expected loss=0.5 P(node) =1
class counts: 5 5
probabilities: 0.500 0.500
left son=2 (6 obs) right son=3 (4 obs)
Primary splits:
Dip < 2.5 to the left, improve=3.3333330, (0 missing)
Test < 1.5 to the left, improve=0.5555556, (0 missing)
Exp < 3.5 to the left, improve=0.5555556, (0 missing)
Node number 2: 6 observations, complexity param=0.1
predicted class=0 expected loss=0.1666667 P(node) =0.6
class counts: 5 1
probabilities: 0.833 0.167
left son=4 (4 obs) right son=5 (2 obs)
Primary splits:
Exp < 4.5 to the left, improve=0.6666667, (0 missing)
Test < 3.5 to the left, improve=0.3333333, (0 missing)
Dip < 1.5 to the right, improve=0.1666667, (0 missing)
Node number 3: 4 observations
predicted class=1 expected loss=0 P(node) =0.4
class counts: 0 4
probabilities: 0.000 1.000
Node number 4: 4 observations
predicted class=0 expected loss=0 P(node) =0.4
class counts: 4 0
probabilities: 1.000 0.000
Node number 5: 2 observations, complexity param=0.1
predicted class=0 expected loss=0.5 P(node) =0.2
class counts: 1 1
probabilities: 0.500 0.500
left son=10 (1 obs) right son=11 (1 obs)
Primary splits:
Test < 2.5 to the left, improve=1, (0 missing)
Node number 10: 1 observations
predicted class=0 expected loss=0 P(node) =0.1
class counts: 1 0
probabilities: 1.000 0.000
Node number 11: 1 observations
predicted class=1 expected loss=0 P(node) =0.1
class counts: 0 1
probabilities: 0.000 1.000 1 2 3 4 5 6 7 8 9 10
0 0 1 0 0 1 1 0 1 1
Levels: 0 1 0 1
1 1 0
2 1 0
3 0 1
4 1 0
5 1 0
6 0 1
7 0 1
8 1 0
9 0 1
10 0 1 [,1] [,2] [,3] [,4] [,5] [,6]
1 1 4 0 1 0 0.4
2 1 4 0 1 0 0.4
3 2 0 1 0 1 0.1
4 1 4 0 1 0 0.4
5 1 4 0 1 0 0.4
6 2 0 4 0 1 0.4
7 2 0 4 0 1 0.4
8 1 1 0 1 0 0.1
9 2 0 4 0 1 0.4
10 2 0 4 0 1 0.4Code
candidates_predict 0 1
1 5 0
2 0 5Code
1 2 3
2 1 1
Classification tree:
rpart(formula = Res ~ Dip + Test + Exp, data = candidates, method = "class",
parms = list(split = "gini"), control = rpart.control(minsplit = 2))
Variables actually used in tree construction:
[1] Dip Exp Test
Root node error: 5/10 = 0.5
n= 10
CP nsplit rel error xerror xstd
1 0.80 0 1.0 2.0 0.00000
2 0.10 1 0.2 0.2 0.18974
3 0.01 3 0.0 0.6 0.28983 [,1] [,2] [,3] [,4] [,5] [,6]
1 1 5 1 0.8333333 0.1666667 0.6
2 1 5 1 0.8333333 0.1666667 0.6
3 1 5 1 0.8333333 0.1666667 0.6
4 1 5 1 0.8333333 0.1666667 0.6
5 1 5 1 0.8333333 0.1666667 0.6
6 2 0 4 0.0000000 1.0000000 0.4
7 2 0 4 0.0000000 1.0000000 0.4
8 1 5 1 0.8333333 0.1666667 0.6
9 2 0 4 0.0000000 1.0000000 0.4
10 2 0 4 0.0000000 1.0000000 0.4References
Breiman, Leo, Jerome H. Friedman, Richard A. Olshen, and Charles J. Stone. 1984. Classification And Regression Trees. 1st ed. Routledge. https://doi.org/10.1201/9781315139470.